 |
Research
at iM2CS
|
|
 |
 |
Research
at iM2CS
|
|
 |
| Our philosophy Modeling involves the development of a mathematical framework to address questions raised from applications. Mathematics provides models’ qualitative behaviors and guides to the design of the most suitable computational method to solve the problem. On the other hand, computations yield quantitative results that can be compared with experimental data, leading to invaluable insights on the phenomena of interest. This process is cyclical and intertwined. The comparison between computational and experimental results often leads to improvements of the mathematical model, whose solutions may predict interesting phenomena not yet observed experimentally. Thus, novel experiments might be designed, leading to substantial progress and advancement in the applied sciences. List of currently active projects:
|
|
|
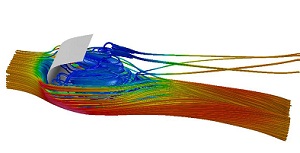 |
Computational fluid and
biofluid dynamics PI: Luoding Zhu Numerical methods are developed and computer simulations are carried out for fundamental mechanical and/or biological processes which involve incompressible viscous fluids and elastic deformable boundaries. There are two major components of the research program: development of numerical methods for fluid-flexible-structure-interaction problems including extension/improvement of the immersed boundary (IB) method, and applications of these methods to problems in life science/ biomedical engineering. Currently work is continuing on: (1) developing a 3D implicit IB method using the lattice Boltzmann approach with applications in problems of biomedical interest -- blood flows with transport and reacting constituents interacting with a compliant vessel wall covered by an endothelial surface layer underlie the initiation and development of atherosclerosis; and the interaction of air flow through the pharyngeal airway with the genioglossal muscle, a process involved in sleep apnea, i.e. a disorder characterized by repetitive collapse of the pharyngeal air way during sleep, and (2) developing novel numerical methods for modeling and simulations of red blood cells interacting with flowing blood using the thin-shell theory and the Navier-Stokes equations. |
|
|
| Computational environmental
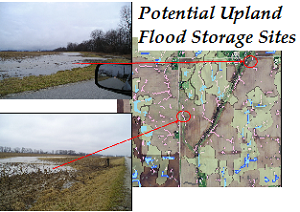
science PIs: Meghna Babbar-Sebens, Snehasis Mukhopadhyay The alteration of the natural hydrologic cycle due to human activities-- such as deforestation, artificial agricultural drainage systems, urbanization, and residential development -- has resulted in loss of multiple ecosystem services (e.g. flood attenuation and water quality control) that were previously provided naturally by various upstream (or upland) landscape features in river basins and watersheds. This research focuses on the restoration of the hydrologic cycle in small watersheds, by designing a system of multiple and distributed upland storage systems all across the landscape. The design process is complex and challenging since it involves selection of multiple sites, multiple scales, structural changes, multiple agronomic practices for agricultural landscapes, and community involvement. To address the complexity of the design process and the acceptability of proposed designs by landowners and local community, the research will develop and integrate computational tools (GIS, simulation, interactive optimization algorithms, etc.) with community participation within the design process. |
 |
|
|
 |
Mathematical and computational
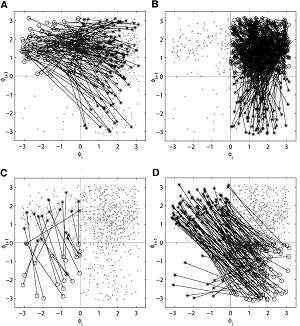
neurosciences PIs: Leonid Rubchinsky, Robert Worth The results of neuroscience research in the last decade have clearlyshown that the knowledge of properties of molecules, genes, neurons and pathways is insufficient to adequately understand many neurological and psychiatric diseases (epilepsy, Parkinson's disease, sleep disorders, memory disorders, schizophrenia, etc.) as well as normal neural processes (motor control, cognitive integration etc.). However, most modern methods in neuroscience research continue to focus on particular elements of neuronal, cellular, or molecular networks, lacking integration of these entities. To integrate the substantial amount of factual knowledge about the elements of the networks, mathematical and computational methods are indispensable. The use of mathematical methods as the basis for the study of the dynamics of systems of neurons is a new and promising direction, which is the focus of our research collaboration. Picture: first-return maps for the |
|
|
| Modeling ocular blood flow and
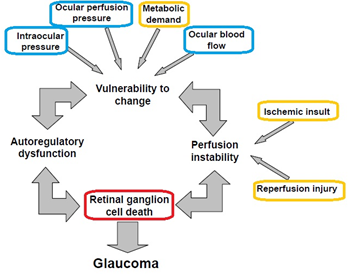
its relation to glaucoma PIs: Giovanna Guidoboni, Alon Harris Glaucoma is a disease in which the optic nerve is damaged, leading toprogressive, irreversible loss of vision. Glaucoma is often associated with increased intraocular pressure (IOP), which is the pressure of the aqueous humor in the eye. Elevated IOP remains the current focus of therapy, but unfortunately many glaucoma patients continue to experience disease progression despite lowered IOP, even to target levels. Clinical observations show that alterations in ocular blood flow play a very important role in the progression of glaucoma. Significant correlations have been found between impaired vascular function and optic nerve damage, but the mechanisms giving rise to these correlations are still unknown. The goal of this project is to investigate the bio-mechanical connections between vascular function and optic nerve damage, in order to gain a better understanding of the risk factors that may be responsible for glaucoma onset and progression. To reach this goal, a variety of mathematical models will be used to describe the different ocular anatomical components, including humors, retina, choroid and sclera. |
 |
|
|
| ABI Innovation: Modeling the
Drosophila Brain with Single-neuron Resolution
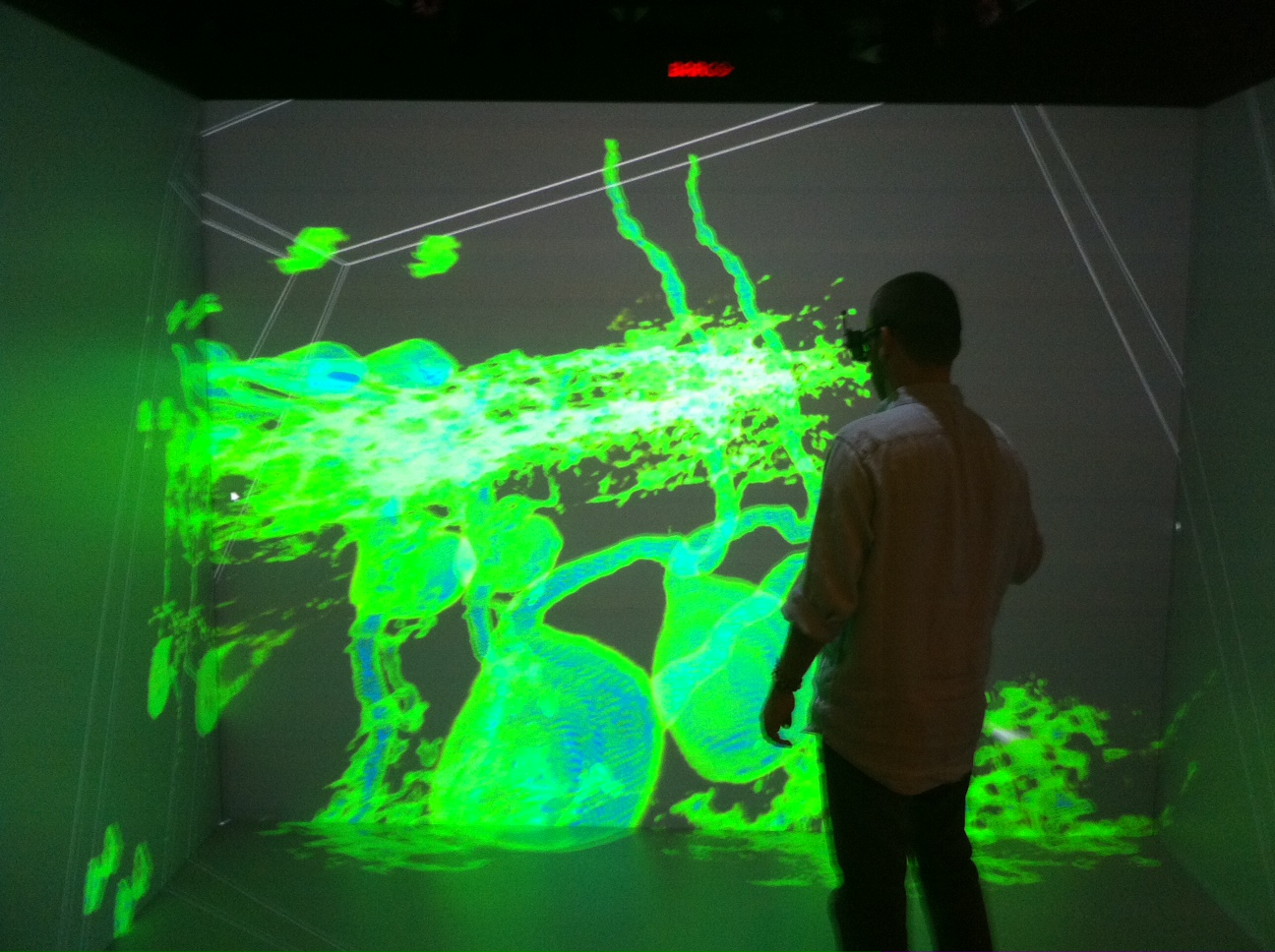
using Computer Vision Methods PI: Gavriil Tsechpenakis The objective of this project is to model the morphology of neurons, how they develop and form synapses in the Central Nervous System (CNS) using computer vision approaches. By exploiting the morphological stereotypes and unparalleled single cell resolution of Drosophila motor neurons, this project will yield the first motor neuron map in the Drosophila CNS, serving as a case study and a step towards modeling the entire brain. The first complete map of a whole brain is that of C. elegans. Its virtue, unchallenged thus far, is resolution down to the level of single visually identified neurons. Because the connectivity of individual neurons ultimately determines brain function, the creation of neural maps at this level in other more complex model organisms is critical for advancing our understanding of brain development. The immediate goal of this project is to create a neural map of a defined population of motor neurons in a healthy Drosophila brain at the single-neuron level. It is intended to be a resource for anyone who wants to ‘navigate’ a model brain in vivo. The long-term goal is to create a complete model brain that will allow for comprehensive analysis of brain development after mutation and the assessment of changes in brain connectivity patterns resulting from drug treatments, disease, or aging and stress. Pictures (click to view in original resolution): top: studying the morphology of individual motor neurons center: tracing and identifying motor neurons in a living brain (courtesy of Akira Chiba, U. of Miami) bottom: virtual navigation in a living Drosophila brain (collaboration with the Advanced Visualization Lab, Indiana University/IUPUI) |
   |
|
|
| Fetal
Alcohol Syndrome Diagnosis
PI: Shiaofen Fang Fetal Alcohol Syndrome (FAS) Disorder is a common alcohol related neurological disorder caused by alcohol exposure during early pregnancy. FAS exhibits characteristic facial features that are often used in FAS diagnosis. Using a 3D laser scanner, we are able to gather detailed 3D facial data (geometry and color) for children of different ages, races and origins with and without FAS. The ability to automatically collect, measure and analyze 3D facial features is very novel in medical diagnosis, and offers unlimited potential for a wide range of future medical solutions. The data analysis process consists of two stages: (1) using currently known FAS related facial features for automatic early diagnosis. This requires 3D interactive visualization, automatic measurements, and 3D surface level computation. (2) 3D feature analysis at both geometry and texture levels, and machine learning based on case studies. Several faculty members in Computer Science are working closely with a group from IU Medical Center as well as several other groups nationwide on a collaborative initiative for FAS diagnosis. |
|
|